FragLite mapping to identify the BRD4 recruitment site of P-TEFb
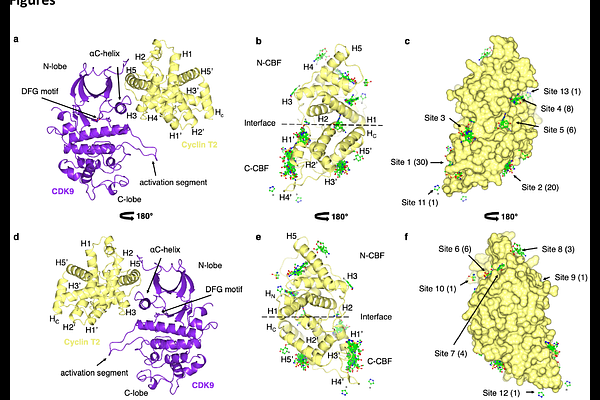
FragLite mapping to identify the BRD4 recruitment site of P-TEFb
Hope, I.; Heath, R.; Basle, A.; Martin, M. P.; Waring, M. J.; Endicott, J. A.; Noble, M. E. M.; Tatum, N. J.
AbstractThe eukaryotic positive transcription elongation factor b (P-TEFb), composed of CDK9 and cyclin T, plays a central role in regulating RNA polymerase II (RNAPII). Phosphorylation of the RNAPII C-terminal domain (CTD) by P-TEFb promotes promoter proximal pause release and enables productive transcriptional elongation across many genes. Cyclin T mediates protein-protein interactions, several of which have been structurally characterised, that help to recruit and fine-tune P-TEFb activity to ensure a tight regulation of transcription. We have previously reported a set of halogenated chemical fragments termed FragLites that can prospectively identify protein interaction sites. Here, we report the FragLite map of cyclin T2, revealing binding sites corresponding to structurally defined cyclin T partners CDK9, AFF4, and HIV-1 Tat. Furthermore, we demonstrate the utility of FragLites in identifying a previously uncharacterised BRD4 binding site. By integrating FragLite clustering with biophysical analyses and AlphaFold3 modelling, we delineate the cyclin T-BRD4 interface. These analyses provide a comprehensive, chemically enriched fragment map highlighting functionally relevant sites to support future probe and modulator development to selectively target P-TEFb.