Enzymatic Ligation Strategy to Enhance Electrospray Ionization Efficiency and Liquid Chromatography-Mass Spectrometry of DNA and RNA Oligonucleotides
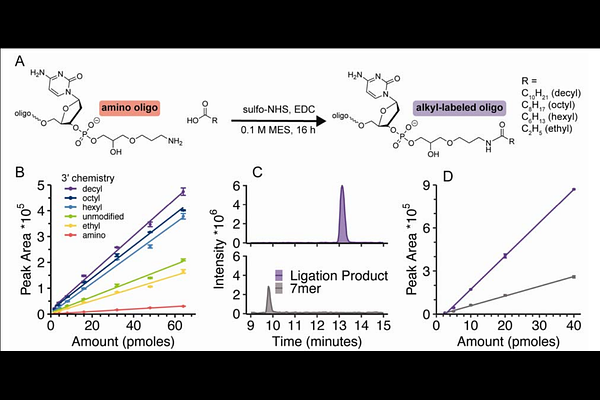
Enzymatic Ligation Strategy to Enhance Electrospray Ionization Efficiency and Liquid Chromatography-Mass Spectrometry of DNA and RNA Oligonucleotides
Sharin, M.; Fitzgerald, N. J.; Kennedy, S. M.; Park, I. G.; Clark, K. D.
AbstractMass spectrometry (MS) is a powerful technique for characterizing modified RNA as it directly sequences and quantifies all mass-altering modifications simultaneously. However, the physicochemical properties of RNA result in poor ionization efficiencies during electrospray ionization, presenting a major barrier to sensitive MS measurements necessary for low abundance RNA samples and RNAs with low modification stoichiometries. Here, we report a ligation-based approach to increase ionization efficiencies of RNA oligonucleotides. We show that short (~5 nt), chemically modified DNA oligonucleotides can be enzymatically ligated to RNA to serve as MS signal enhancers. Among a series of signal enhancers appended with various alkyl and alkylimidazolium functional groups, we found that decyl-functionalized derivatives improved MS sensitivity by ~15-fold compared to unlabeled oligonucleotide. When ligated to RNA standards, the decyl-modified signal enhancer increased MS signals 2-4-fold with the additional benefit of improved retention during liquid chromatography (LC) separations without ion pairing agents. To apply the ligation-based approach to RNase T1 digests of longer RNAs, a multi-step enzymatic approach was optimized to maximize ligation efficiencies. We then ligated signal enhancers to a yeast transfer RNA (tRNA) digest and observed increased MS signals for numerous sequence-informative digestion products. Importantly, the sequences of RNA oligonucleotides ligated to signal enhancers were readily determined by tandem mass spectrometry with collision-induced dissociation. This ligation-based strategy for enhancing LC-MS/MS characterization of RNA creates opportunities to measure low abundance RNA samples and their modifications.