Comparison of the Distribution of Fitness Effects Across Primates
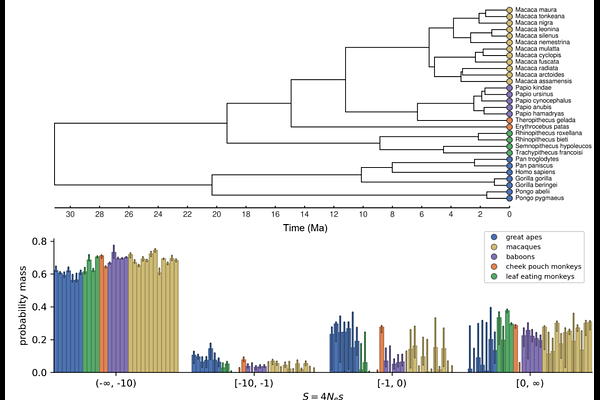
Comparison of the Distribution of Fitness Effects Across Primates
Sendrowski, J.; Pedersen, B. M.; Bergman, J.; Pankratov, V.; Bataillon, T.
AbstractThe distribution of fitness effects (DFE) of mutations is a key determinant of both the efficacy of natural selection and the genetic load of populations. It also provides an indirect summary of the underlying fitness landscape, so the extent to which the DFE is conserved across species can offer insights into the invariance of these landscapes. Here, we infer the DFE of amino acid--changing mutations in 38 catarrhine primates using site frequency spectrum (SFS)--based methods. Our results are consistent with the nearly neutral theory, with interspecific differences primarily driven by variation in effective population size (Ne) and thus the efficacy of selection. Specifically, a one-order-of-magnitude increase in Ne is associated with an approximately 10% increase in the fraction of strongly deleterious mutations. Although DFE estimates exhibit clade-level clustering, this pattern largely reflects shared Ne rather than clade-specific effects, and in unscaled units, deleterious DFEs are broadly similar across species. These conclusions are robust to the choice of DFE parameterization, phylogenetic regression framework, and ancestral misidentification. Extending the analysis to non-additive dominance effects further shows that dominance is only weakly identifiable from the SFS and has minimal impact on comparative DFE inference. Finally, simulations demonstrate that demographic correction via nuisance parameters enables robust inference across a range of demographic scenarios.