A foundation AI model enhances electron microscopy image analysis
A foundation AI model enhances electron microscopy image analysis
Du, M.; Wang, Y.; Xie, L.; Deng, G.; Guo, J.; Han, B.; Chen, Z.-H.; Rui, C.; Han, J.; Chen, Y.; Zhao, Y.; Cao, R.; Wang, F.; Li, K.; Wang, Y.; He, Y.; feng, x.
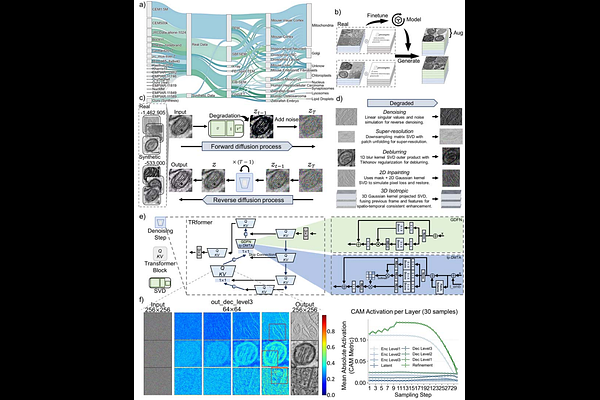
AbstractElectron microscopy (EM) and advanced volume EM have extensive applications in deciphering cellular ultrastructures for life sciences. However, quality issues of EM images significantly impede the precise analysis and discovery of nanoscale biological structures. Here, we present an unsupervised Foundation Model for Five Tasks (DF5T) model for image enhancement through denoising, deblurring, super-resolution, two-dimensional (2D) inpainting, and three-dimensional (3D) isotropic restoration. DF5T was trained on an extensive dataset of multi-source membrane-bound organelle EM images, consisting of a total of over 2.25 million images. On unseen data, DF5T outperformed the existing state-of-the-art models in all five tasks. Furthermore, DF5T substantially compensates for missing three-dimensional structural information and reveals significant ultrastructural alterations in organelle geometry following chemical treatment. We demonstrate that DF5T significantly enhances electron microscopy images quality by improving the accuracy of downstream organelle segmentation and 3D structural restoration, thereby contributing to future advances in biological research.