Interrogating the Escherichia coli epitranscriptome via CRISPR interference and Nanopore native RNA sequencing
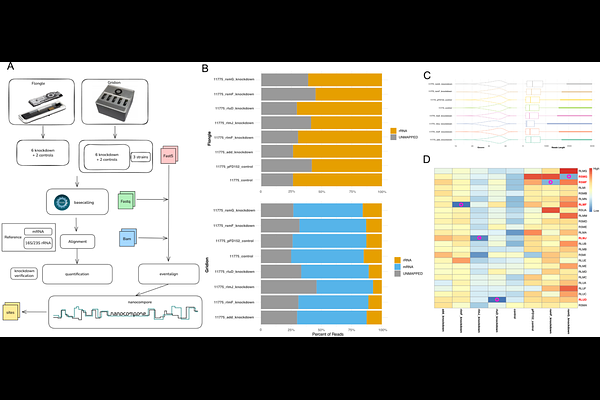
Interrogating the Escherichia coli epitranscriptome via CRISPR interference and Nanopore native RNA sequencing
Pitt, M. E.; Zhang, J.; Nguyen, A. N. T.; Hall, M. B.; Jebeli, L.; Featherstone, L. A.; Myers, G. S. A.; Scott, N.; Coin, L. J. M.
AbstractEpitranscriptomics has recently gained significant momentum due to technological advances and translational applications, however, studies on bacterial RNA modifications remain limited. Bacterial RNA remains notoriously prone to degradation and methodologies to investigate the epitranscriptome are challenging. Prior research has shown RNA modifications modulate antimicrobial resistance, virulence and pathogenicity. This research employed CRISPR interference to knock down five known Escherichia coli rRNA modification genes (rlmF, rlmJ, rluD, rsmF and rsmG) in three E. coli strains. These isolates underwent growth curves, proteome analysis and native RNA sequencing. CRISPRi adequately silenced the majority of RNA modification genes in E. coli (>80% reduction). Significant growth delays were associated with rlmF, rsmF and rsmG repression. Unique protein pathways corresponding with RNA modification loss were found for rlmJ (TreB, XylF), rluD (CysH, HycB, PutP, TrpB), rsmF (EvgA) and rsmG (OppC). Known rRNA modification sites for rluD ({Psi}) and rsmG (m7G) were detected from analysis of nanopore electrical signal, however, only a weak signal was apparent for m6A (rlmF, rlmJ) and m5C (rsmF) modifications. The inhibition of rRNA modifications resulted in mRNA modification changes including for genes ompC, cspC, dbhA, dbhB and secY. Our work provides an approach for unravelling the epitranscriptome of E. coli and gain insight into its functional role.