An autonomous system for multi-objective continuous evolution at scale
An autonomous system for multi-objective continuous evolution at scale
Boileau, R. M.; Golas, S. M.; Ma, Q.; Jiang, B.; a, A.; Jia, M.; Ilieva, N.; Baydush, A.; Fu, H.; Chory, E. J.
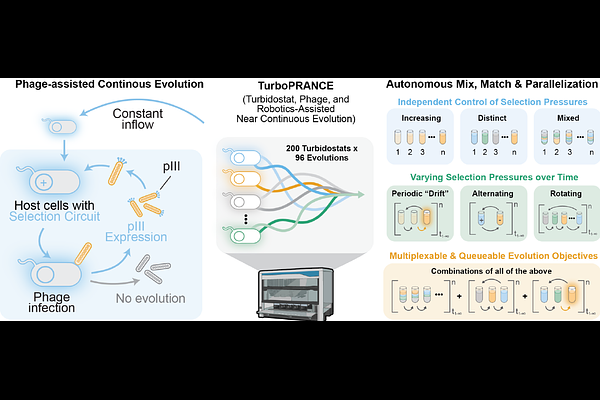
AbstractNatural evolution is high-dimensional; organisms adapt to many pressures at once, across substrates, environments, and genetic backgrounds. Yet most directed evolution methods flatten this landscape to a single selection axis, hiding tradeoffs, and limiting what can be learned. Phage-assisted continuous evolution (PACE) is uniquely suited for multivariate selection because horizontal gene transfer couples genotype to propagation and allows the same phage lineage to traverse different selection environments. In practice, implementing this at scale has been prohibitive because each selection demands its own host culture, and every culture must be held for days to weeks within a narrow, infectable density window using continuously responsive bioreactors. In this work, TurboPRANCE is presented as an open-source, queueable robotic platform that integrates ~200 independently controlled turbidostats with 96 parallel PACE lagoons under closed-loop control. Each turbidostat operates as a fully separate unit that can be equilibrated and initiated on its own schedule, enabling asynchronous starts and sustained operation without intervention. Automated media formulation, programmable dosing, on-deck sterilization, and adaptive scheduling coordinate growth control with the changing needs of the robotic workflow, dynamically adjusting dilution and transfer timing around formulation, sampling, and handling steps to keep each culture at consistent infectable densities despite unpredictable method demands. Cultures can be multiplexed and titrated into lagoons at defined ratios, swapped in and out on a schedule, or kept fully separate across experiments, creating a combinatorial space of selection pressures and programs that is effectively unbounded. Additionally, to enable high-throughput evolutionary tracking that scales with TurboPRANCE, Nanopore long-read sequencing was combined with DeepVariant, a deep learning-based variant caller, enabling population-level tracking of evolving variants. The result is a system that generates high-resolution time-resolvable evolutionary trajectories and large parallel datasets spanning diverse selection regimes, yielding dense, multivariate training data to map and engineer complex fitness landscapes at scale.