Quantitative prediction of nonsense-mediated mRNA decay across human genes by genomic language model and large-scale mutational scanning
Quantitative prediction of nonsense-mediated mRNA decay across human genes by genomic language model and large-scale mutational scanning
Veiner, M.; Toledano, I.; Palou-Marquez, G.; Lehner, B.; Supek, F.
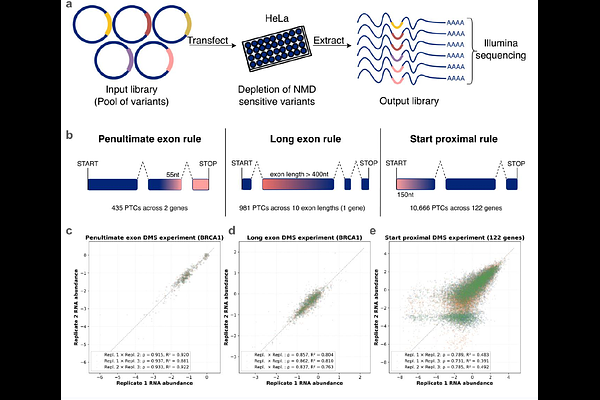
AbstractThe molecular consequences of protein truncating variants depend strongly on whether their transcripts are eliminated by nonsense-mediated mRNA decay (NMD), yet NMD is still predicted largely from a small set of binary positional rules. How individual premature termination codons (PTCs) engage NMD across genes and transcript contexts therefore remains incompletely resolved. Here we integrated endogenous allele-specific PTC expression from large-scale genomic data, mRNA language-model prediction and high-throughput mutational scanning to revisit the rules that govern mammalian NMD. Using allele-specific expression measurements from large human cohorts, we trained NMDetective-AI on ~14,000 somatic PTCs, and after testing it on ~1,800 germline PTCs, found that it improves on previous models, with its accuracy approaching the reproducibility of the underlying measurements. We then generated experimental maps of NMD with deep mutational scanning, including ~450 PTCs nearby the 50-nt penultimate exon boundary, ~950 engineered PTCs across 9 exon lengths to resolve long-exon escape, and ~11k PTCs across 139 genes to quantify and refine start-proximal evasion. Our results support the positional logic of mammalian NMD yet show that this logic is implemented quantitatively: the classical rules resolve into graded, gene-dependent response curves whose boundaries are shaped by transcript architecture and modulated by local sequence context. Applying this framework to population, disease and cancer datasets further identifies genes in which NMD is predicted to aggravate or ameliorate the effects of truncating variants, providing a basis for variant interpretation and for prioritizing NMD-directed therapies.