Structural variants contribute substantially to complex trait heritability
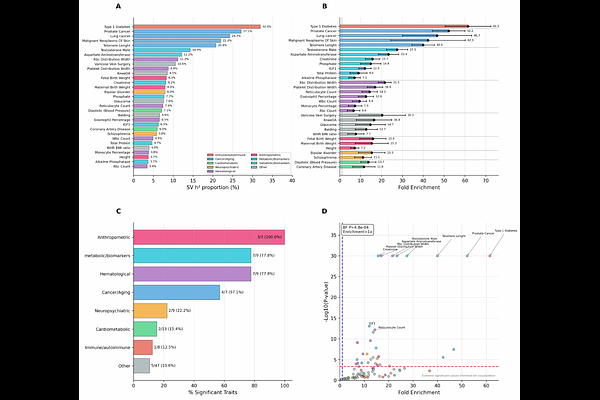
Structural variants contribute substantially to complex trait heritability
Nguyen, D. T.; Shadrin, A. A.; Parker, N.; Fuhrer, J.; Vo, N. S.; Dale, A.; Andreassen, O.; Frei, O.
AbstractDespite accumulating evidence that structural variants (SVs) exert disproportionate functional effects, their genome-wide contribution to complex trait heritability has not yet been systematically quantified. Here, we introduce MiXeR-SV, a tool that integrates long-read-derived SVs from existing reference catalogs with genome-wide association summary statistics to quantify SV heritability and enrichment. Applying MiXeR-SV to 105 complex traits, we identify 31 traits with significant enrichment (Bonferroni-corrected P < 4.8*10-4;), with SVs explaining up to 32% of total heritability, despite comprising only 0.6% of the analyzed variants. Enrichment is most extensive in hematological, metabolic/biomarker, and cancer traits, trait-specific in neuropsychiatric and cardiometabolic phenotypes, and observed across all anthropometric traits, albeit with modest effect sizes, consistent with their highly polygenic architecture. These findings are robust across two independent SV reference panels built from complementary long-read and graph-based variant catalogs, with 93.5% of significant traits consistent between technologies. Furthermore, incorporating SV architecture into heritability models consistently increases total heritability estimates in enriched phenotypes, with SV heritability proportions correlating with the gap between SNP-based and twin heritability estimates across traits. Cross-ancestry replication in Biobank Japan confirms enrichment patterns. Our quantification of SV contributions to genetic architectures has significant implications for genetic prediction and fine-mapping of human complex traits and common diseases.