A rapid, sensitive, and quantitative high plex biomarker digital detection platform enabled by Hypercoding
A rapid, sensitive, and quantitative high plex biomarker digital detection platform enabled by Hypercoding
Bathina, M.; Blum, A. P.; Brodin, J.; DeBuono, N.; Fu, Y.; Lu, B.; Naticchia, M. R.; Ortiz, D.; Richards, A.; Rozieres, C. d.; Schowalter, R.; Shultzaberger, S.; Snow, S.; Tanner, S.; Trejo, C. L.; Ward, S.; LeCoultre, R.; Read, K.; Sathe, S.; Schlegel, C.; Schlegel, I.; Shaner, S.; Tsay, J.; Weir, J.; Wong, K. M.; Abi-Samra, K.; Alldredge, J.; Anderson, P.; Bailey, J.; Bollig, C.; Bonnardel, J.; Bru, A.; Chan, A. C. S.; Chang, T.; DeBerg, L.; Doorn, J. v.; Driscoll, P.; Duarte, T.; Esparza, A.; Frerichs, D.; Gautherot, A.; Held, L.; Hendricks, G.; Holst, G.; Iwamoto, K.; Jimenez, H.; Khandan,
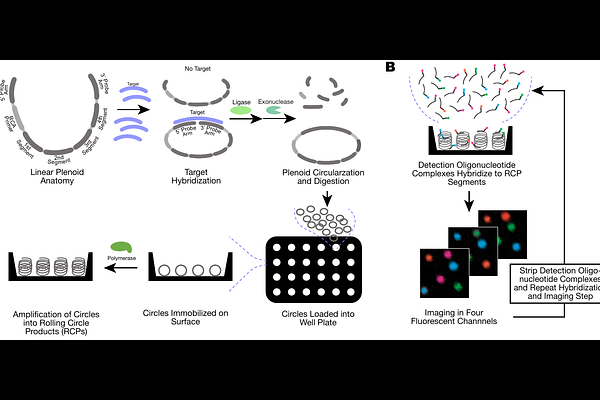
AbstractLow-cost, multiplexed, and automated assays are needed to make omic technologies more broadly accessible in clinical, research, and commercial settings. We present HypercodingTM, a scalable technology for detection and quantitation of multi-omic targets. Drawing from data reliability methods in the telecommunications field, Hypercoding uses fluorescent signals from hybridization with an error-correcting code to enable detection of high-plexity targets from biological samples, such as human DNA. In the presence of the target, a linear DNA construct is circularized, immobilized, and amplified to enable single-molecule detection of a target via rapid readout cycles within a 96-well plate. We demonstrate capability for >10,000 code plexity and accurate (98.7%) genotyping of 209 pharmacogenomic variants. Furthermore, we show computation of copy number variation with whole chromosome and sub-gene resolution, as well as quantitation of target abundance down to 10 fM sensitivity with a dynamic range of up to 10 logs.