Systematic errors in enzymatic conversion limit cell-free DNA methylation specificity
Voice is AI-generated
Connected to paperThis paper is a preprint and has not been certified by peer review
Systematic errors in enzymatic conversion limit cell-free DNA methylation specificity
Loyfer, N.; Magenheim, J.; Darwish, A.; Isaac, S.; Ganbat, J.; Babikir, H.; Jhutty, A.; Wan, J.; Bayes-Genis, A.; Revuelta-Lopez, E.; Eden, A.; Solanki, R.; Dor, Y.; Kaplan, T.
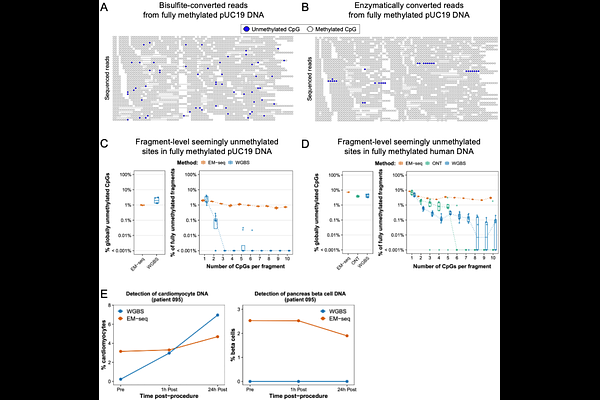
AbstractEnzymatic methylation sequencing (EM-seq) converts unmethylated cytosines to uracils while preserving DNA integrity, making it attractive for liquid biopsies. Here we report a reproducible fragment-level over-conversion error in EM-seq, which is not observed in bisulfite-based conversion or in Oxford Nanopore sequencing. While bisulfite and nanopore errors occur at sporadic CpGs, EM-seq generates molecules that appear fully unmethylated, introducing a false-positive background signal that severely limits deconvolution specificity in cfDNA analysis.