The DoGA Consortium Atlas of Canine Enhancers and Promoters Across Tissues and Development
The DoGA Consortium Atlas of Canine Enhancers and Promoters Across Tissues and Development
Takan, I.; Hortenhuber, M.; Salokorpi, N.; Bokhari, R.; Araujo, C.; Aljelaify, R.; Quintero, I.; Ezer, S.; Mottaghitalab, F.; Raman, A.; Ross, F.; DoGA Consortium, ; Jokinen, T. S.; Syrja, P.; Bannasch, D.; Iivanainen, A.; Hytönen, M. K.; Kere, J.; Lohi, H.; Daub, C. O.
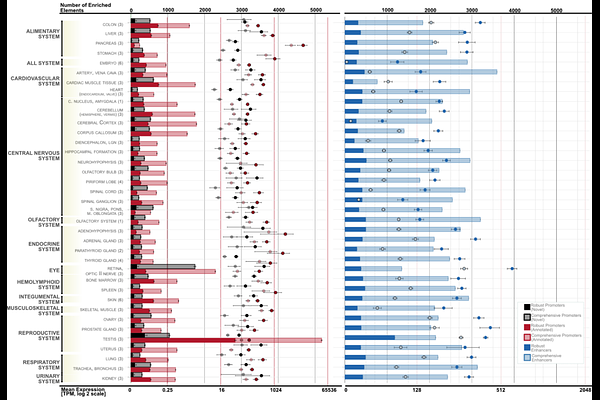
AbstractThe domestic dog is a powerful genetic model for complex traits, disease and behaviours relevant to human biology, yet the canine genome still lacks a systematic transcription-defined atlas of regulatory elements. Existing resources have improved annotation but largely infer regulatory activity from chromatin features rather than directly mapping transcription initiation at promoters and enhancers. Here, we address this gap using 114 CAGE-seq libraries spanning 56 tissues and developmental stages from 9 dogs. We identify 68,446 promoters and 46,661 active enhancers, and define their tissue-enriched activity across the canine body. Regulatory programmes are organized around shared transcription factor motif infrastructures, with prominent enrichment of KLF family motifs across tissues. The cerebellum emerges as a major regulatory hub, showing the highest density of enhancer-promoter interactions among brain regions. During embryogenesis, enhancer activity shifts from early neurodevelopmental patterning programmes, including OTX2 and ZIC1/4, towards regulators of functional maturation and synaptic organization, including AQP4 and NLGN3. Comparative analyses revealed 69 enhancers with highly similar regulatory structures linked to potential orthologous genes in dog and human. Together, these data establish a tissue-resolved transcription-based atlas of the canine regulatory genome that advances functional annotation and improves interpretation of non-coding variation in comparative and disease genomics.