Multiscale confidence quantification for virtual spatial transcriptomics with UTOPIA
Multiscale confidence quantification for virtual spatial transcriptomics with UTOPIA
Jin, K.; Chen, Z.; Yu, X.; Yuan, M.; Schroeder, A.; Dumoulin, B.; Liu, Y.; Wang, L.; Park, J. H.; Hwang, T. H.; Susztak, K.; Ren, Z.; Zhang, N.; Li, M.
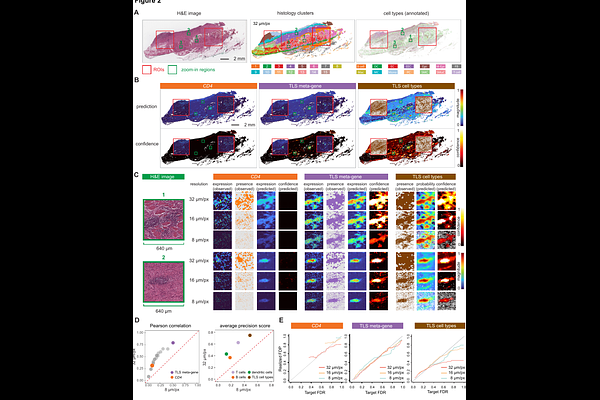
AbstractVirtual spatial transcriptomics (ST) methods predict gene expression or cell types from histology images, extending molecular readouts beyond the limited regions or samples directly measured by ST platforms. However, the statistical reliability of these predictions remains unclear. Here, we present UTOPIA, a model-agnostic framework for multiscale confidence quantification in virtual ST. UTOPIA assigns statistically calibrated confidence scores to predictions across spatial resolutions and biological granularities, ranging from single genes to metagenes and from specific cell types to broader cell classes. UTOPIA controls false discovery rates for detecting genes, metagenes, or cell types while accounting for local tissue context. We show that prediction confidence depends critically on both spatial resolution and biological granularity, with reliable inference often emerging only at coarser, biologically meaningful scales. Across multiple ST platforms and in both in-sample and out-of-sample settings, UTOPIA enhances interpretability, prevents false biological conclusions, and enables more trustworthy downstream analyses of virtual ST.