Multiscale Physical Effects of CpG Methylation on DNA Mechanics, Nucleosome Wrapping, and Chromatin Condensates
Multiscale Physical Effects of CpG Methylation on DNA Mechanics, Nucleosome Wrapping, and Chromatin Condensates
Daris, L. C.; Rizvi, A.; Singh, A. A.; Hutchings, J.; Lamers, L. A.; Montabana, E. A.; Kimanius, D.; Banerjee, P. R.; Eeftens, J. M.; Sanulli, S.
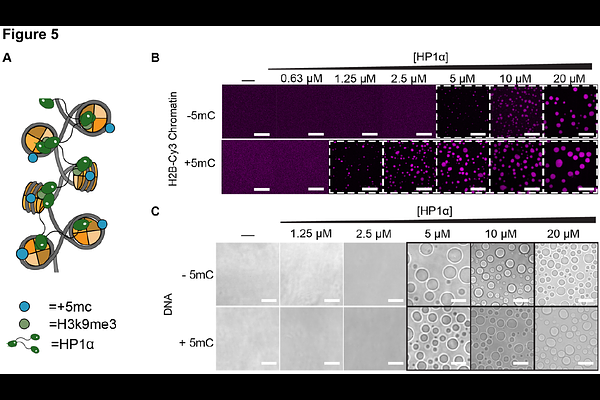
AbstractDNA methylation at CpG dinucleotides is a central epigenetic modification linked to transcriptional repression and heterochromatin formation, yet its physical impact on chromatin organization remains incompletely understood. Here, we integrate single-molecule force spectroscopy, FRAP, and nanorheology to quantify how CpG methylation alters chromatin mechanics across scales. Optical tweezer-based measurements show that CpG methylation modestly increases DNA contour and persistence lengths, indicating a slightly extended and stiffer DNA helix. When assembled into nucleosomes, CpG methylation weakens both outer- and inner turn wrapping, reducing unwrapping forces and per-nucleosome unwrapping energy. On a higher-order assembly scale, although CpG methylated and unmethylated chromatin fibers display similar thresholds for salt-induced phase separation, methylated condensates exhibit markedly reduced internal mobility: FRAP reveals slower chromatin exchange, nanorheology shows increased viscoelasticity and longer relaxation times. Finally, CpG methylation enhances HP1a-chromatin interactions, lowering the HP1a concentration required for condensation, despite no change in HP1a binding to naked DNA. Together, these results show that CpG methylation remodels chromatin mechanics by tuning DNA rigidity, nucleosome wrapping energetics, and condensate material properties, providing a multiscale physical framework for how DNA methylation contributes to heterochromatin.