Investigating dietary microRNA stability and function using a transgenic milk model with unique microRNA sequences
Investigating dietary microRNA stability and function using a transgenic milk model with unique microRNA sequences
Husseini, Z.; Majeau, N.; Fliss, I.; Benmousa, A.
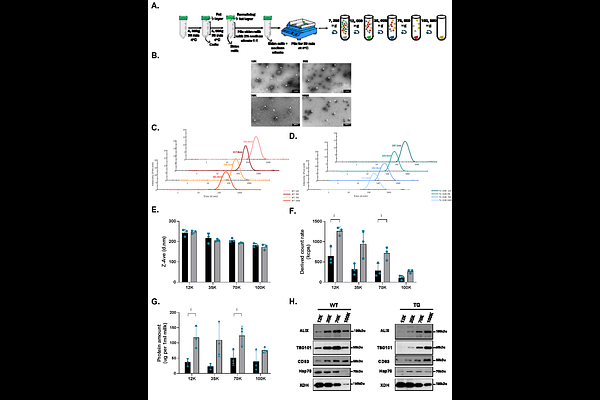
AbstractMilk microRNAs are believed to play gene regulatory functions in the consumer's cells. Milk from different species is enriched in microRNAs predicted to influence immunity, metabolism, and intestinal homeostasis. For milk microRNAs to regulate gene expression in the consumer, they must survive digestion and be present at sufficient levels to influence intestinal cells and potentially beyond-intestinal cells. Milk microRNAs are proposed to be protected from degradation through their association with milk extracellular vesicles (EVs), which might also deliver them to cells. Studies on milk microRNA oral transfer and tissue bioavailability are limited by interspecies sequence homology, making it difficult to distinguish endogenous from exogenous microRNAs. Here, we used a transgenic (TG) cow model expressing four unique microRNA sequences (AmiRs) in its milk to study their association with milk EVs, their resistance to in vitro digestion, and AmiR uptake and regulatory activity in vitro. We confirmed the presence of the four milk EV populations in raw wild-type (WT) and TG cow milk, similar to those previously reported in commercial (pasteurized) cow milk, and confirmed their association with AmiRs and classical milk microRNAs. AmiRs showed differential resistance to simulated adult digestion. In vitro uptake studies showed a modest gene regulatory effect of AmiRs in Caco-2 cells incubated with TG milk EVs. The intent of using this model was to perform in vitro analysis which could lay the groundwork for later in vivo bioavailability studies, taking advantage of the uniqueness of the AmiRs sequences and bypassing the limitation of microRNA sequence homology.