SWANS: A highly configurable analysis pipeline for single-cell and single-nuclei RNA-sequencing data
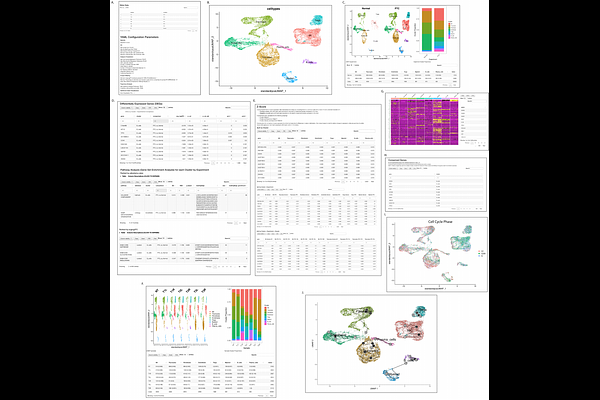
SWANS: A highly configurable analysis pipeline for single-cell and single-nuclei RNA-sequencing data
Beigel, K.; Wafula, E.; Mitchelle, D. V.; Pastor, S. J.; Gong, M.; Heuckeroth, R.; Ricarte-Filho, J. C.; Franco, A. T.; Reichenberger, E. R.
AbstractBackground: Single-cell RNA sequencing (scRNA-seq) is a powerful technique that enables the analysis of gene expression at the individual cell level. Bioinformatic tools for scRNA-seq data analysis have many different options throughout the typical scRNA-seq workflow (normalization, integration, annotation, clustering, and visualization), and the choice of method(s) and parameter(s) at each stage can impact results. Results: Here, we introduce SWANS (v2.0), a configurable analysis pipeline that, in a single run, can employ multiple analysis methods, resolutions, and modifiable parameters. The resulting clustering arrangements, differential gene expression results, and other quantitative measurements can be dynamically visualized and compared in a Shiny interactive report to assist in choosing a single analysis schema for annotation and downstream analysis. Once a final approach is chosen, SWANS will perform differential gene expression (DGE) analysis based on experimental conditions and gene set enrichment analysis (GSEA) in addition to creating reports that display figures and interactive tables, quality control metrics, and benchmarking information. SWANS uses Snakemake as a workflow manager, Cell Ranger for alignment and gene expression quantification, Seurat for single cell data analysis, and additional single cell R packages for quality control and downstream single cell analysis. Conclusion: SWANS is a tailorable pipeline that provides options for quality control, dimensionality reduction, clustering, differential gene expression analysis, gene set enrichment analysis, and trajectory analysis. Additionally, SWANS generates a series of reports that facilitate sharing large volumes of complex data in a clear and concise manner with other investigators.