STRmie-HD enables interruption-aware HTT repeat genotyping and somatic mosaicism profiling across sequencing platforms
STRmie-HD enables interruption-aware HTT repeat genotyping and somatic mosaicism profiling across sequencing platforms
Napoli, A.; Liorni, N.; Biagini, T.; Giovannetti, A.; Squitieri, A.; Miele, L.; Urbani, A.; Caputo, V.; Gasbarrini, A.; Squitieri, F.; Mazza, T.
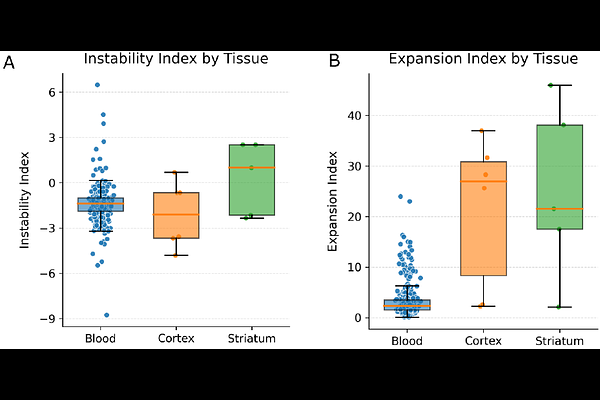
AbstractShort tandem repeat expansions in exon 1 of the HTT gene drive Huntington's disease (HD) pathogenesis, with disease onset and progression heavily influenced by somatic mosaicism and sequence interruptions. While sequencing technologies enable repeat sizing, many computational tools lack the resolution to capture subtle interruption motifs and allele-specific somatic variation. We present STRmie-HD, an alignment-free, de novo framework for interruption-aware genotyping and quantitative profiling of somatic mosaicism at single-read resolution. The tool parses individual reads to quantify uninterrupted CAG tract length, CCG repeat content, and critical interruption variants, including Loss of Interruption (LOI) and Duplication of Interruption (DOI). Validated across Illumina, PacBio SMRT, and Oxford Nanopore platforms, STRmie-HD demonstrates high concordance with reference genotypes and superior sensitivity in identifying rare interruption patterns that conventional tools often overlook. Furthermore, it implements somatic mosaicism metrics to characterize repeat dynamics and successfully distinguishes a higher somatic expansion burden in brain tissues than in peripheral blood. STRmie-HD offers a comprehensive and extensible solution for high-resolution molecular characterization of HTT variation, providing a robust framework for patient stratification and genetic research in HD.