DNA methylation signatures of mismatch repair-deficient colorectal cancer
DNA methylation signatures of mismatch repair-deficient colorectal cancer
Ward, R.; Endicott, M.; Mallabar-Rimmer, B.; Burrage, J.; Sherwood, K.; Huang, Q.; Ward, J. C.; Thorn, S.; Woolley, C.; Wood, S.; Dempster, E.; Green, H. D.; Tomlinson, I.; Webster, A. P.
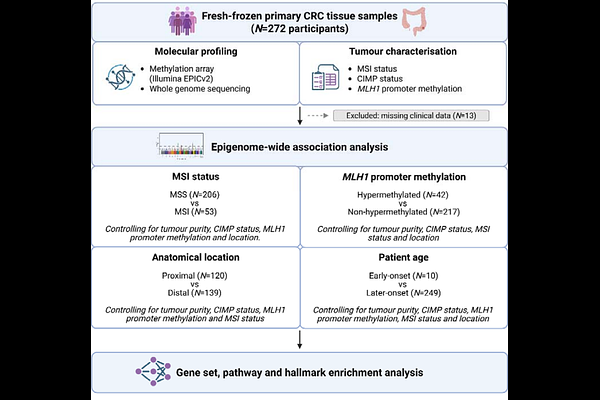
AbstractBackground: Colorectal cancer (CRC) is a molecularly heterogeneous disease shaped by both genetic and epigenetic alterations. Approximately 15% of CRCs display widespread CpG island hypermethylation, known as the CpG Island Methylator Phenotype (CIMP). CIMP-high (CIMP-H) tumours frequently exhibit MLH1 promoter hypermethylation, leading to mismatch repair deficiency (MMRd) and microsatellite instability (MSI). However, DNA methylation patterns associated with MSI, independent of CIMP and MLH1 silencing, and the influence of clinical variables such as anatomical location and patient age on the CRC methylome remain poorly characterised. Methods: We performed epigenome-wide DNA methylation profiling of 259 primary CRC tissue samples using the Illumina EPICv2 array, comparing differential methylation between MSI and microsatellite stable (MSS) CRC, adjusting for tumour purity, MLH1 promoter methylation, CIMP status, and anatomical location, to account for known confounders. We further evaluated the independent effects of anatomical location and patient age on global methylation patterns. Results: Epigenome-wide differential methylation between MSS and MSI CRC was dominated by MLH1 promoter hypermethylation. After adjusting for MLH1 hypermethylation and CIMP status, we identified a distinct set of 656 CpG sites associated with MMRd independent of MLH1 silencing. These included hypermethylation at LRP6, GSK3{beta}, and CDK12, implicating altered WNT signalling and transcriptional regulation pathways. Comparison of MSI subgroups revealed the co-occurrence of MLH1 hypermethylation with promoter hypermethylation at TXNRD1. Anatomical location showed a strong independent effect on methylation patterns, while we observed only modest effects of patient age on the CRC methylome after adjustment for confounders. Conclusions: We identified a distinct methylation profile distinguishing MSS and MSI CRC, including MLH1-independent markers of MMRd, as well as novel differentially methylated loci within MSI subgroups. We further showed that anatomical location has a strong independent impact on the CRC methylome. Together, these findings refine the molecular characterisation of CRC and highlight potential epigenetic markers that could inform patient stratification and precision oncology.