EvoMut: A Computational Framework for Engineering Oxidative Stability in Proteins
EvoMut: A Computational Framework for Engineering Oxidative Stability in Proteins
Arab, S. S.; Lewis, N. E.
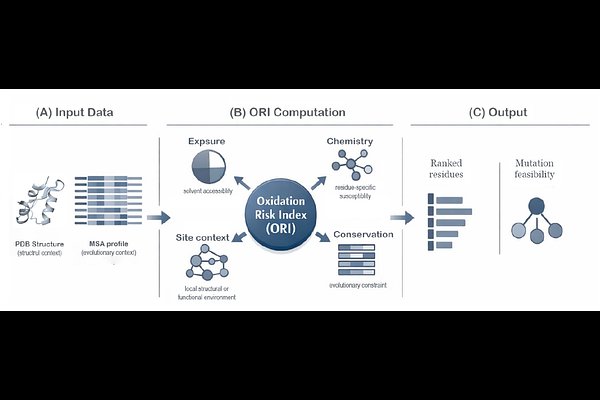
AbstractAmino acid oxidation is a major cause of protein instability and loss of function in therapeutic and industrial settings. Although methionine, cysteine, tyrosine, and tryptophan residues are widely recognized as oxidation-prone, only a subset of such residues are dominant functional hotspots, and not all are suitable targets for mutation. Identifying these vulnerable yet engineerable sites remains a major challenge. Here, we present EvoMut, a residue-level analytical framework for evaluating both oxidative vulnerability and mutation feasibility. EvoMut estimates oxidation risk by integrating structural features, local functional context, intrinsic chemical susceptibility, and evolutionary conservation. A central feature of the framework is the explicit separation of oxidation risk from mutation feasibility: candidate substitutions are evaluated only after high-risk residues are identified and ranked by evolutionary substitution patterns. Application of EvoMut to multiple proteins, and evaluation with experimental data, showed that oxidation-prone residues differ markedly in their engineering potential. EvoMut distinguishes residues that are both oxidation-sensitive and evolutionarily permissive from those that are chemically vulnerable but functionally constrained. By providing residue-level mechanistic insight, EvoMut offers a practical framework for the rational design of oxidation-resistant proteins. EvoMut is freely available as a web server at https://evomut.org.