snputils: A High-Performance Python Library for Genetic Variation and Population Structure
snputils: A High-Performance Python Library for Genetic Variation and Population Structure
Bonet, D.; Comajoan Cara, M.; Barrabes, M.; Smeriglio, R.; Agrawal, D.; Aounallah, K.; Geleta, M.; Dominguez Mantes, A.; Thomassin, C.; Shanks, C.; Huang, E. C.; Franquesa Mones, M.; Luis, A.; Saurina, J.; Perera, M.; Lopez, C.; Sabat, B. O.; Abante, J.; Moreno-Grau, S.; Mas Montserrat, D.; Ioannidis, A. G.
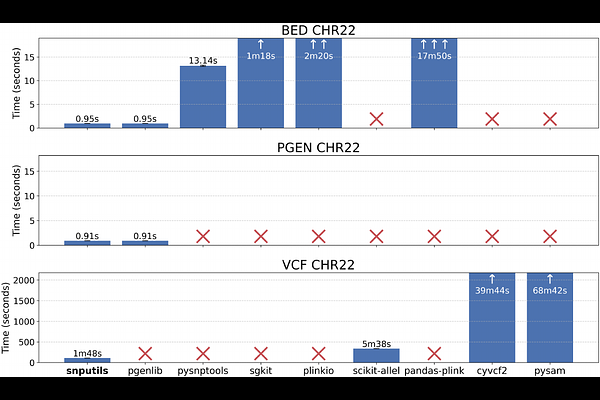
AbstractThe increasing size and resolution of genomic and population genetic datasets offer unprecedented opportunities to study population structure and uncover the genetic basis of complex traits and diseases. The collection of existing analytical tools, however, is characterized by format incompatibilities, limited functionality, and computational inefficiencies, forcing researchers to construct fragile pipelines that chain together fragmented command-line utilities and ad hoc scripts. These are difficult to maintain, scale, and reproduce. To address such limitations, we present snputils, a Python library that unifies high-performance I/O, transformation, and analysis of genotype, ancestry, and phenotypic information within a single framework suitable for biobank-scale research. The library provides efficient tools for essential operations, including querying, cleaning, merging, and statistical analysis. In addition, it offers classical population genetic statistics with optional ancestry-specific masking. An identity-by-descent module supports reading of multiple formats, filtering and ancestry-restricted segment trimming for relatedness and demographic inference. snputils also incorporates ancestry-masking and multi-array functionalities for dimensionality reduction methods, as well as efficient implementations of admixture simulation, admixture mapping, and advanced visualization capabilities. With support for the most commonly used file formats, snputils integrates smoothly with existing tools and clinical databases. At the same time, its modular and optimized design reduces technical overhead, facilitating reproducible workflows that accelerate discoveries in population genetics, genomic research, and precision medicine. Benchmarking demonstrates a significant reduction in genotype data loading speed compared to existing Python libraries. The open-source library is available at https://github.com/AI-sandbox/snputils, with full documentation and tutorials at https://snputils.org.