LLPSight: enhancing prediction of LLPS-driving proteins using machine learning and protein Language Models
LLPSight: enhancing prediction of LLPS-driving proteins using machine learning and protein Language Models
GONAY, V.; VITALE, R.; STEGMAYER, G.; Dunne, M. P.; KAJAVA, A. V.
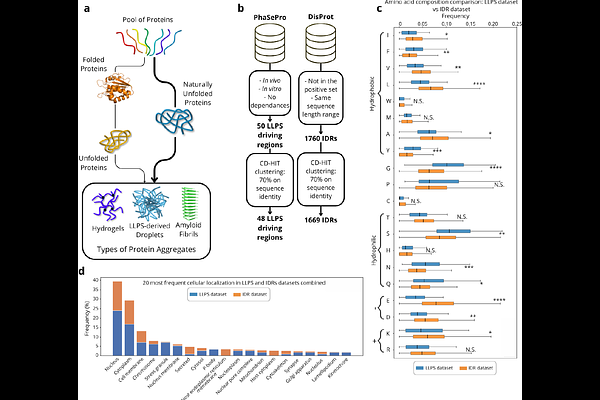
AbstractIn eukaryotic cells, essential functions are often confined within organelles enclosed by lipid membranes. Increasing evidence, however, highlights the role of membrane-less organelles (MLOs), formed through liquid-liquid phase separation (LLPS). MLO assemblies are typically initiated by >>driver>> proteins, which form a scaffold to recruit additional >>client>> molecules. By leveraging expanding MLO datasets and modern machine learning approaches, we developed LLPSight, an ML-based predictor of LLPS-driving proteins. The model was trained using rigorously curated datasets: a positive set of proteins experimentally confirmed to drive LLPS in vivo and a negative set of soluble, unstructured proteins not associated with LLPS. For the features, we employed a cutting-edge approach using embeddings from protein Language Models. LLPSight achieves the highest F1 score (0.885) among existing tools, enabling more efficient discovery of new LLPS drivers eagerly awaited by researchers for experimental validation. An additional key feature of LLPSight is its ability to perform proteome-wide analyses; application to the human proteome yielded promising targets. LLPSight can be obtained from authors upon request.